| Protein or domain name | Size | Number of residues with known Φ-values | Predicted amyloidogenic regions | Residues with experimental Φ-values larger than 0.50 |
| Spliceosomal protein U1A | 96 | 10 | 44(0.60) | |
| Ribosomal protein S6 | 97 | 10 | 6(0.52) | |
| B-domain of staphylococcal protein A | 60 | 15 | ||
| Colicin E9 immunity protein | 86 | 19 | ||
| Barnase | 110 | 19 | ||
| Chymotrypsin inhibitor (CI2) | 64 | 34 | ||
| Ig repeat 27 of titin | 89 | 22 | ||
| IgG binding domain of streptococcal protein L | 64 | 38 | ||
| Villin 14T domain 1 | 126 | 18 | ||
| 10th FN3 module of fibronectin | 94 | 16 | 28–35; | |
| DNA-binding protein Sso7d | 64 | 15 | ||
| Barstar | 89 | 22 | ||
| Human muscle acylphosphatase | 98 | 17 | ||
| FK-506 binding protein (FKBP12) | 107 | 22 | ||
| SH3 domain of src tyrosine kinase transforming protein | 64 | 26 | ||
| Colicin E7 immunity protein | 87 | 19 | ||
| CheY | 128 | 9 | ||
| IgG binding domain of streptococcal protein G | 56 | 20 | ||
| SH3 domain of α-spectrin | 62 | 13 | ||
| 3rd Fn3 repeat of tenascin (short form) | 90 | 25 | ||
aIf several Φ-values are known for a single residue (several different mutations were made), the Φ-values were averaged.
bFor these proteins, amyloidogenic regions are revealed only with window size of five residues. The size of the sliding window for the rest proteins was seven residues.
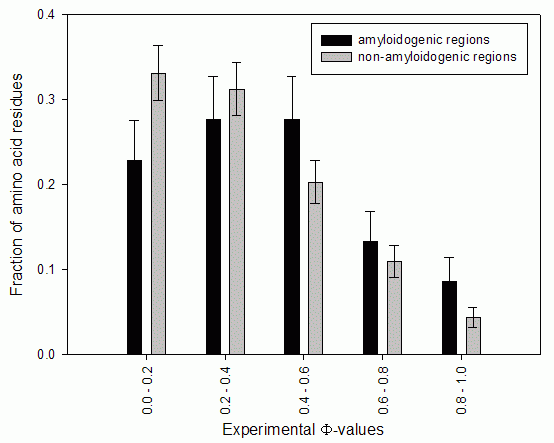
The fraction of all known experimental Φ-values in the predicted amyloidogenic regions is 25%, the fraction of large Φ-values (>0.5) within amyloidogenic fragments is 33%, such Φ-values are shown in bold in Table.

Experimentally measured and theoretically predicted Φ-values.